Phylogenetics
| Part of a series on |
| Evolutionary biology |
|---|
Key topics
|
Processes and outcomes
|
Natural history
|
History of evolutionary theory
|
Fields and applications
|
Social implications
|
|
In biology, phylogenetics /ˌfaɪloʊdʒəˈnɛtɪks, -lə-/[1][2] (Greek: φυλή, φῦλον – phylé, phylon = tribe, clan, race + γενετικός – genetikós = origin, source, birth)[3] is the study of the evolutionary history and relationships among individuals or groups of organisms (e.g. species, or populations). These relationships are discovered through phylogenetic inference methods that evaluate observed heritable traits, such as DNA sequences or morphology under a model of evolution of these traits. The result of these analyses is a phylogeny (also known as a phylogenetic tree) – a diagrammatic hypothesis about the history of the evolutionary relationships of a group of organisms.[4] The tips of a phylogenetic tree can be living organisms or fossils, and represent the "end", or the present, in an evolutionary lineage. Phylogenetic analyses have become central to understanding biodiversity, evolution, ecology, and genomes.
Taxonomy is the identification, naming and classification of organisms. It is usually richly informed by phylogenetics, but remains a methodologically and logically distinct discipline.[5] The degree to which taxonomies depend on phylogenies (or classification depends on evolutionary development) differs depending on the school of taxonomy: phenetics ignores phylogeny altogether, trying to represent the similarity between organisms instead; cladistics (phylogenetic systematics) tries to reproduce phylogeny in its classification without loss of information; evolutionary taxonomy tries to find a compromise between them.
Contents
1 Construction of a phylogenetic tree
2 History
2.1 Ernst Haeckel's recapitulation theory
2.2 Timeline of key events
3 See also
4 References
5 Bibliography
6 External links
Construction of a phylogenetic tree
Usual methods of phylogenetic inference involve computational approaches implementing the optimality criteria and methods of parsimony, maximum likelihood (ML), and MCMC-based Bayesian inference. All these depend upon an implicit or explicit mathematical model describing the evolution of characters observed.
Phenetics, popular in the mid-20th century but now largely obsolete, used distance matrix-based methods to construct trees based on overall similarity in morphology or other observable traits (i.e. in the phenotype, not the DNA), which was often assumed to approximate phylogenetic relationships.
Prior to 1990, phylogenetic inferences were generally presented as narrative scenarios. Such methods are often ambiguous and lack explicit criteria for evaluating alternative hypotheses.[6][7][8]
History
The term "phylogeny" derives from the German Phylogenie, introduced by Haeckel in 1866,[9] and the Darwinian approach to classification became known as the "phyletic" approach.[10]
Ernst Haeckel's recapitulation theory
During the late 19th century, Ernst Haeckel's recapitulation theory, or "biogenetic fundamental law", was widely accepted. It was often expressed as "ontogeny recapitulates phylogeny", i.e. the development of a single organism during its lifetime, from germ to adult, successively mirrors the adult stages of successive ancestors of the species to which it belongs. But this theory has long been rejected.[11][12] Instead, ontogeny evolves – the phylogenetic history of a species cannot be read directly from its ontogeny, as Haeckel thought would be possible, but characters from ontogeny can be (and have been) used as data for phylogenetic analyses; the more closely related two species are, the more apomorphies their embryos share.
Timeline of key events
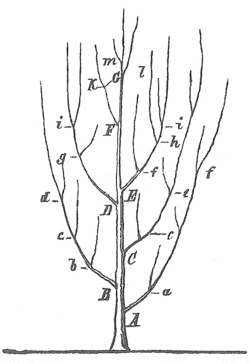
Branching tree diagram from Heinrich Georg Bronn's work (1858)

Phylogenetic tree suggested by Haeckel (1866)
- 14th century, lex parsimoniae (parsimony principle), William of Ockam, English philosopher, theologian, and Franciscan friar, but the idea actually goes back to Aristotle, precursor concept
- 1763, Bayesian probability, Rev. Thomas Bayes,[13] precursor concept
- 18th century, Pierre Simon (Marquis de Laplace), perhaps first to use ML (maximum likelihood), precursor concept
- 1809, evolutionary theory, Philosophie Zoologique, Jean-Baptiste de Lamarck, precursor concept, foreshadowed in the 17th century and 18th century by Voltaire, Descartes, and Leibniz, with Leibniz even proposing evolutionary changes to account for observed gaps suggesting that many species had become extinct, others transformed, and different species that share common traits may have at one time been a single race,[14] also foreshadowed by some early Greek philosophers such as Anaximander in the 6th century BC and the atomists of the 5th century BC, who proposed rudimentary theories of evolution[15]
- 1837, Darwin's notebooks show an evolutionary tree[16]
- 1843, distinction between homology and analogy (the latter now referred to as homoplasy), Richard Owen, precursor concept
- 1858, Paleontologist Heinrich Georg Bronn (1800–1862) published a hypothetical tree to illustrating the paleontological "arrival" of new, similar species following the extinction of an older species. Bronn did not propose a mechanism responsible for such phenomena, precursor concept.[17]
- 1858, elaboration of evolutionary theory, Darwin and Wallace,[18] also in Origin of Species by Darwin the following year, precursor concept
- 1866, Ernst Haeckel, first publishes his phylogeny-based evolutionary tree, precursor concept
- 1893, Dollo's Law of Character State Irreversibility,[19] precursor concept
- 1912, ML recommended, analyzed, and popularized by Ronald Fisher, precursor concept
- 1921, Tillyard uses term "phylogenetic" and distinguishes between archaic and specialized characters in his classification system[20]
- 1940, term "clade" coined by Lucien Cuénot
- 1949, Jackknife resampling, Maurice Quenouille (foreshadowed in '46 by Mahalanobis and extended in '58 by Tukey), precursor concept
- 1950, Willi Hennig's classic formalization[21]
- 1952, William Wagner's groundplan divergence method[22]
- 1953, "cladogenesis" coined[23]
- 1960, "cladistic" coined by Cain and Harrison[24]
- 1963, first attempt to use ML (maximum likelihood) for phylogenetics, Edwards and Cavalli-Sforza[25]
- 1965
- Camin-Sokal parsimony, first parsimony (optimization) criterion and first computer program/algorithm for cladistic analysis both by Camin and Sokal[26]
- character compatibility method, also called clique analysis, introduced independently by Camin and Sokal (loc. cit.) and E. O. Wilson[27]
- Camin-Sokal parsimony, first parsimony (optimization) criterion and first computer program/algorithm for cladistic analysis both by Camin and Sokal[26]
- 1966
- English translation of Hennig[28]
- "cladistics" and "cladogram" coined (Webster's, loc. cit.)
- English translation of Hennig[28]
- 1969
- dynamic and successive weighting, James Farris[29]
- Wagner parsimony, Kluge and Farris[30]
- CI (consistency index), Kluge and Farris[30]
- introduction of pairwise compatibility for clique analysis, Le Quesne[31]
- dynamic and successive weighting, James Farris[29]
- 1970, Wagner parsimony generalized by Farris[32]
- 1971
- first successful application of ML to phylogenetics (for protein sequences), Neyman[33]
- Fitch parsimony, Fitch[34]
- NNI (nearest neighbour interchange), first branch-swapping search strategy, developed independently by Robinson[35] and Moore et al.
- ME (minimum evolution), Kidd and Sgaramella-Zonta[36] (it is unclear if this is the pairwise distance method or related to ML as Edwards and Cavalli-Sforza call ML "minimum evolution")
- first successful application of ML to phylogenetics (for protein sequences), Neyman[33]
- 1972, Adams consensus, Adams[37]
- 1976, prefix system for ranks, Farris[38]
- 1977, Dollo parsimony, Farris[39]
- 1979
- Nelson consensus, Nelson[40]
- MAST (maximum agreement subtree)((GAS)greatest agreement subtree), a consensus method, Gordon [41]
- bootstrap, Bradley Efron, precursor concept[42]
- Nelson consensus, Nelson[40]
- 1980, PHYLIP, first software package for phylogenetic analysis, Felsenstein
- 1981
- majority consensus, Margush and MacMorris[43]
- strict consensus, Sokal and Rohlf[44]
- first computationally efficient ML algorithm, Felsenstein[45]
- majority consensus, Margush and MacMorris[43]
- 1982
- PHYSIS, Mikevich and Farris
- branch and bound, Hendy and Penny[46]
- 1985
- first cladistic analysis of eukaryotes based on combined phenotypic and genotypic evidence Diana Lipscomb[47]
- first issue of Cladistics
- first phylogenetic application of bootstrap, Felsenstein[48]
- first phylogenetic application of jackknife, Scott Lanyon[49]
- first cladistic analysis of eukaryotes based on combined phenotypic and genotypic evidence Diana Lipscomb[47]
- 1986, MacClade, Maddison and Maddison
- 1987, neighbor-joining method Saitou and Nei[50]
- 1988, Hennig86 (version 1.5), Farris
- Bremer support (decay index), Bremer[51]
- Bremer support (decay index), Bremer[51]
- 1989
- RI (retention index), RCI (rescaled consistency index), Farris[52]
- HER (homoplasy excess ratio), Archie[53]
- RI (retention index), RCI (rescaled consistency index), Farris[52]
- 1990
- combinable components (semi-strict) consensus, Bremer[54]
- SPR (subtree pruning and regrafting), TBR (tree bisection and reconnection), Swofford and Olsen[55]
- combinable components (semi-strict) consensus, Bremer[54]
- 1991
- DDI (data decisiveness index), Goloboff[56][57]
- first cladistic analysis of eukaryotes based only on phenotypic evidence, Lipscomb
- DDI (data decisiveness index), Goloboff[56][57]
- 1993, implied weighting Goloboff[58]
- 1994, reduced consensus: RCC (reduced cladistic consensus) for rooted trees, Wilkinson[59]
- 1995, reduced consensus RPC (reduced partition consensus) for unrooted trees, Wilkinson[60]
- 1996, first working methods for BI (Bayesian Inference)independently developed by Li,[61] Mau,[62] and Rannala and Yang[63] and all using MCMC (Markov chain-Monte Carlo)
- 1998, TNT (Tree Analysis Using New Technology), Goloboff, Farris, and Nixon
- 1999, Winclada, Nixon
- 2003, symmetrical resampling, Goloboff[64]
See also
 Phylogenetics portal
Phylogenetics portal
- Angiosperm Phylogeny Group
- Bauplan
- Bioinformatics
- Biomathematics
- Coalescent theory
- EDGE of Existence programme
- Evolutionary taxonomy
- Joe Felsenstein
- Language family
- Maximum parsimony
- Microbial phylogenetics
- Molecular phylogeny
- Noogenesis
- Ontogeny (psychoanalysis)
- PhyloCode
- Phylodynamics
- Phylogenesis
- Phylogenetic comparative methods
- Phylogenetic network
- Phylogenetic nomenclature
- Phylogenetic tree viewers
- Phylogenetics software
- Phylogenomics
- Phylogeny (psychoanalysis)
- Phylogeography
- Systematics
References
^ "phylogenetic". Dictionary.com Unabridged. Random House..mw-parser-output cite.citation{font-style:inherit}.mw-parser-output q{quotes:"""""""'""'"}.mw-parser-output code.cs1-code{color:inherit;background:inherit;border:inherit;padding:inherit}.mw-parser-output .cs1-lock-free a{background:url("//upload.wikimedia.org/wikipedia/commons/thumb/6/65/Lock-green.svg/9px-Lock-green.svg.png")no-repeat;background-position:right .1em center}.mw-parser-output .cs1-lock-limited a,.mw-parser-output .cs1-lock-registration a{background:url("//upload.wikimedia.org/wikipedia/commons/thumb/d/d6/Lock-gray-alt-2.svg/9px-Lock-gray-alt-2.svg.png")no-repeat;background-position:right .1em center}.mw-parser-output .cs1-lock-subscription a{background:url("//upload.wikimedia.org/wikipedia/commons/thumb/a/aa/Lock-red-alt-2.svg/9px-Lock-red-alt-2.svg.png")no-repeat;background-position:right .1em center}.mw-parser-output .cs1-subscription,.mw-parser-output .cs1-registration{color:#555}.mw-parser-output .cs1-subscription span,.mw-parser-output .cs1-registration span{border-bottom:1px dotted;cursor:help}.mw-parser-output .cs1-hidden-error{display:none;font-size:100%}.mw-parser-output .cs1-visible-error{font-size:100%}.mw-parser-output .cs1-subscription,.mw-parser-output .cs1-registration,.mw-parser-output .cs1-format{font-size:95%}.mw-parser-output .cs1-kern-left,.mw-parser-output .cs1-kern-wl-left{padding-left:0.2em}.mw-parser-output .cs1-kern-right,.mw-parser-output .cs1-kern-wl-right{padding-right:0.2em}
^ "Phylogenetic". Merriam-Webster Dictionary.
^ Liddell, Henry George; Scott, Robert; Jones, Henry Stuart (1968). A Greek-English lexicon (9 ed.). Oxford: Clarendon Press. p. 1961.
^ "phylogeny". Biology online. Retrieved 2013-02-15.
^ Edwards AWF; Cavalli-Sforza LL (1964). "Reconstruction of evolutionary trees". In Heywood, Vernon Hilton; McNeill, J. Phenetic and Phylogenetic Classification. pp. 67–76. OCLC 733025912.Phylogenetics is the branch of life science concerned with the analysis of molecular sequencing data to study evolutionary relationships among groups of organisms.
^ Richard C. Brusca & Gary J. Brusca (2003). Invertebrates (2nd ed.). Sunderland, Massachusetts: Sinauer Associates.
ISBN 978-0-87893-097-5.
^ Bock, W. J. (2004). Explanations in systematics. Pp. 49–56. In Williams, D. M. and Forey, P. L. (eds) Milestones in Systematics. London: Systematics Association Special Volume Series 67. CRC Press, Boca Raton, Florida.
^ Auyang, Sunny Y. (1998). Narratives and Theories in Natural History. In: Foundations of complex-system theories: in economics, evolutionary biology, and statistical physics. Cambridge, U.K.; New York: Cambridge University Press.[page needed]
^ Harper, Douglas (2010). "Phylogeny". Online Etymology Dictionary. Retrieved March 18, 2013.
^ Stuessy 2009.
^ Blechschmidt, Erich (1977) The Beginnings of Human Life. Springer-Verlag Inc., p. 32: "The so-called basic law of biogenetics is wrong. No buts or ifs can mitigate this fact. It is not even a tiny bit correct or correct in a different form, making it valid in a certain percentage. It is totally wrong."
^ Ehrlich, Paul; Richard Holm; Dennis Parnell (1963) The Process of Evolution. New York: McGraw–Hill, p. 66: "Its shortcomings have been almost universally pointed out by modern authors, but the idea still has a prominent place in biological mythology. The resemblance of early vertebrate embryos is readily explained without resort to mysterious forces compelling each individual to reclimb its phylogenetic tree."
^ Bayes, Mr; Price, Mr (1763). "An Essay towards Solving a Problem in the Doctrine of Chances. By the Late Rev. Mr. Bayes, F. R. S. Communicated by Mr. Price, in a Letter to John Canton, A. M. F. R. S". Philosophical Transactions of the Royal Society of London. 53: 370–418. doi:10.1098/rstl.1763.0053.
^ Strickberger, Monroe. 1996. Evolution, 2nd. ed. Jones & Bartlett.[page needed]
^ The Theory of Evolution, Teaching Company course, Lecture 1
^ Darwin's Tree of Life Archived 13 March 2014 at the Wayback Machine.
^ Archibald, J. David (2008). "Edward Hitchcock's Pre-Darwinian (1840) 'Tree of Life'". Journal of the History of Biology. 42 (3): 561–92. CiteSeerX 10.1.1.688.7842. doi:10.1007/s10739-008-9163-y. PMID 20027787.
^ Darwin, Charles; Wallace, Alfred (1858). "On the Tendency of Species to form Varieties; and on the Perpetuation of Varieties and Species by Natural Means of Selection". Journal of the Proceedings of the Linnean Society of London. Zoology. 3 (9): 45–62. doi:10.1111/j.1096-3642.1858.tb02500.x.
^ Dollo, Louis. 1893. Les lois de l'évolution. Bull. Soc. Belge Géol. Paléont. Hydrol. 7: 164–66.
^ Tillyard, R. J (2012). "A New Classification of the Order Perlaria". The Canadian Entomologist. 53 (2): 35–43. doi:10.4039/Ent5335-2.
^ Hennig, Willi (1950). Grundzüge einer Theorie der Phylogenetischen Systematik [Basic features of a theory of phylogenetic systematics] (in German). Berlin: Deutscher Zentralverlag. OCLC 12126814.
[page needed]
^ Wagner, Warren Herbert (1952). "The fern genus Diellia: structure, affinities, and taxonomy". University of California Publications in Botany. 26 (1–6): 1–212. OCLC 4228844.
^ Webster's 9th New Collegiate Dictionary
^ Cain, A. J; Harrison, G. A (2009). "Phyletic Weighting". Proceedings of the Zoological Society of London. 135 (1): 1–31. doi:10.1111/j.1469-7998.1960.tb05828.x.
^ "The reconstruction of evolution" in "Abstracts of Papers". Annals of Human Genetics. 27 (1): 103–5. 1963. doi:10.1111/j.1469-1809.1963.tb00786.x.
^ Camin, Joseph H; Sokal, Robert R (1965). "A Method for Deducing Branching Sequences in Phylogeny". Evolution. 19 (3): 311–26. doi:10.1111/j.1558-5646.1965.tb01722.x.
^ Wilson, Edward O (1965). "A Consistency Test for Phylogenies Based on Contemporaneous Species". Systematic Zoology. 14 (3): 214–20. doi:10.2307/2411550. JSTOR 2411550.
^ Hennig. W. (1966). Phylogenetic systematics. Illinois University Press, Urbana.[page needed]
^ Farris, James S (1969). "A Successive Approximations Approach to Character Weighting". Systematic Zoology. 18 (4): 374–85. doi:10.2307/2412182. JSTOR 2412182.
^ ab Kluge, A. G; Farris, J. S (1969). "Quantitative Phyletics and the Evolution of Anurans". Systematic Biology. 18 (1): 1–32. doi:10.1093/sysbio/18.1.1.
^ Quesne, Walter J. Le (1969). "A Method of Selection of Characters in Numerical Taxonomy". Systematic Zoology. 18 (2): 201–205. doi:10.2307/2412604. JSTOR 2412604.
^ Farris, J. S (1970). "Methods for Computing Wagner Trees". Systematic Biology. 19: 83–92. doi:10.1093/sysbio/19.1.83.
^ Neyman, J. (1971). Molecular studies: A source of novel statistical problems. In: Gupta S. S., Yackel J. (eds), Statistical Decision Theory and Related Topics, pp. 1–27. Academic Press, New York.
^ Fitch, W. M (1971). "Toward Defining the Course of Evolution: Minimum Change for a Specific Tree Topology". Systematic Biology. 20 (4): 406–16. doi:10.1093/sysbio/20.4.406. JSTOR 2412116.
^ Robinson, D.F (1971). "Comparison of labeled trees with valency three". Journal of Combinatorial Theory, Series B. 11 (2): 105–19. doi:10.1016/0095-8956(71)90020-7.
^ Kidd, K. K; Sgaramella-Zonta, L. A (1971). "Phylogenetic analysis: Concepts and methods". American Journal of Human Genetics. 23 (3): 235–52. PMC 1706731. PMID 5089842.
^ Adams, E. N (1972). "Consensus Techniques and the Comparison of Taxonomic Trees". Systematic Biology. 21 (4): 390–397. doi:10.1093/sysbio/21.4.390.
^ Farris, James S (1976). "Phylogenetic Classification of Fossils with Recent Species". Systematic Zoology. 25 (3): 271–282. doi:10.2307/2412495. JSTOR 2412495.
^ Farris, J. S (1977). "Phylogenetic Analysis Under Dollo's Law". Systematic Biology. 26: 77–88. doi:10.1093/sysbio/26.1.77.
^ Nelson, G (1979). "Cladistic Analysis and Synthesis: Principles and Definitions, with a Historical Note on Adanson's Familles Des Plantes (1763-1764)". Systematic Biology. 28: 1–21. doi:10.1093/sysbio/28.1.1.
^ Gordon, A. D (1979). "A Measure of the Agreement between Rankings". Biometrika. 66 (1): 7–15. doi:10.2307/2335236. JSTOR 2335236.
^ Efron B. (1979). Bootstrap methods: another look at the jackknife. Ann. Stat. 7: 1–26.
^ Margush, T; McMorris, F (1981). "Consensus-trees". Bulletin of Mathematical Biology. 43 (2): 239. doi:10.1016/S0092-8240(81)90019-7.
^ Sokal, Robert R; Rohlf, F. James (1981). "Taxonomic Congruence in the Leptopodomorpha Re-Examined". Systematic Zoology. 30 (3): 309. doi:10.2307/2413252. JSTOR 2413252.
^ Felsenstein, Joseph (1981). "Evolutionary trees from DNA sequences: A maximum likelihood approach". Journal of Molecular Evolution. 17 (6): 368–76. doi:10.1007/BF01734359. PMID 7288891.
^ Hendy, M.D; Penny, David (1982). "Branch and bound algorithms to determine minimal evolutionary trees". Mathematical Biosciences. 59 (2): 277. doi:10.1016/0025-5564(82)90027-X.
^ Lipscomb, Diana. 1985. The Eukaryotic Kingdoms. Cladistics 1: 127–40.
^ Felsenstein J. (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783–791.
^ Lanyon, S. M (1985). "Detecting Internal Inconsistencies in Distance Data". Systematic Biology. 34 (4): 397–403. CiteSeerX 10.1.1.1000.3956. doi:10.1093/sysbio/34.4.397.
^ Saitou, N.; Nei, M. (1987). "The neighbor-joining method: A new method for reconstructing phylogenetic trees". Molecular Biology and Evolution. 4 (4): 406–25. doi:10.1093/oxfordjournals.molbev.a040454. PMID 3447015.
^ Bremer, Kåre (1988). "The Limits of Amino Acid Sequence Data in Angiosperm Phylogenetic Reconstruction". Evolution. 42 (4): 795–803. doi:10.1111/j.1558-5646.1988.tb02497.x. PMID 28563878.
^ Farris, James S (1989). "The Retention Index and the Rescaled Consistency Index". Cladistics. 5 (4): 417–419. doi:10.1111/j.1096-0031.1989.tb00573.x.
^ Archie, James W (1989). "Homoplasy Excess Ratios: New Indices for Measuring Levels of Homoplasy in Phylogenetic Systematics and a Critique of the Consistency Index". Systematic Zoology. 38 (3): 253–269. doi:10.2307/2992286. JSTOR 2992286.
^ Bremer, Kåre (1990). "Combinable Component Consensus". Cladistics. 6 (4): 369–372. doi:10.1111/j.1096-0031.1990.tb00551.x.
^ D. L. Swofford and G. J. Olsen. 1990. Phylogeny reconstruction. In D. M. Hillis and G. Moritz (eds.), Molecular Systematics, pages 411–501. Sinauer Associates, Sunderland, Mass.
^ Goloboff, Pablo A (1991). "Homoplasy and the Choice Among Cladograms". Cladistics. 7 (3): 215–232. doi:10.1111/j.1096-0031.1991.tb00035.x.
^ Goloboff, Pablo A (1991). "Random Data, Homoplasy and Information". Cladistics. 7 (4): 395–406. doi:10.1111/j.1096-0031.1991.tb00046.x.
^ Goloboff, Pablo A (1993). "Estimating Character Weights During Tree Search". Cladistics. 9: 83–91. doi:10.1111/j.1096-0031.1993.tb00209.x.
^ Wilkinson, M (1994). "Common Cladistic Information and its Consensus Representation: Reduced Adams and Reduced Cladistic Consensus Trees and Profiles". Systematic Biology. 43 (3): 343–368. doi:10.1093/sysbio/43.3.343.
^ Wilkinson, Mark (1995). "More on Reduced Consensus Methods". Systematic Biology. 44 (3): 435–439. doi:10.2307/2413604. JSTOR 2413604.
^ Li, Shuying; Pearl, Dennis K; Doss, Hani (2000). "Phylogenetic Tree Construction Using Markov Chain Monte Carlo". Journal of the American Statistical Association. 95 (450): 493. CiteSeerX 10.1.1.40.4461. doi:10.1080/01621459.2000.10474227. JSTOR 2669394.
^ Mau, Bob; Newton, Michael A; Larget, Bret (1999). "Bayesian Phylogenetic Inference via Markov Chain Monte Carlo Methods". Biometrics. 55 (1): 1–12. CiteSeerX 10.1.1.139.498. doi:10.1111/j.0006-341X.1999.00001.x. JSTOR 2533889. PMID 11318142.
^ Rannala, Bruce; Yang, Ziheng (1996). "Probability distribution of molecular evolutionary trees: A new method of phylogenetic inference". Journal of Molecular Evolution. 43 (3): 304–11. doi:10.1007/BF02338839. PMID 8703097.
^ Goloboff, P (2003). "Improvements to resampling measures of group support". Cladistics. 19 (4): 324–32. doi:10.1016/S0748-3007(03)00060-4.
Bibliography
.mw-parser-output .refbegin{font-size:90%;margin-bottom:0.5em}.mw-parser-output .refbegin-hanging-indents>ul{list-style-type:none;margin-left:0}.mw-parser-output .refbegin-hanging-indents>ul>li,.mw-parser-output .refbegin-hanging-indents>dl>dd{margin-left:0;padding-left:3.2em;text-indent:-3.2em;list-style:none}.mw-parser-output .refbegin-100{font-size:100%}
Schuh, Randall T.; Brower, Andrew V.Z. (2009). Biological Systematics: principles and applications (2nd ed.). Ithaca: Comstock Pub. Associates/Cornell University Press. ISBN 978-0-8014-4799-0. OCLC 312728177.
Forster, Peter; Renfrew, Colin, eds. (2006). Phylogenetic Methods and the Prehistory of Languages. McDonald Institute Press, University of Cambridge. ISBN 978-1-902937-33-5. OCLC 69733654.
Baum, David A.; Smith, Stacey D. (2013). Tree Thinking: an introduction to phylogenetic biology. Greenwood Village, CO: Roberts and Company. ISBN 978-1-936221-16-5. OCLC 767565978.
Stuessy, Tod F. (2009). Plant Taxonomy: The Systematic Evaluation of Comparative Data. Columbia University Press. ISBN 978-0-231-14712-5. Retrieved 6 February 2014.
External links
| Look up phylogenetics in Wiktionary, the free dictionary. |